This is a guest post by Class-Thido Pfaff We here present the BEFdata R package as part of the rOpenSci project. It is an API package that combines the strengths of the BEFdata portal in handling small, complex datasets with the powerful statics package R. The portal itself is free software as well and can be found here.
Many US federal agencies are now running app competitions to highlight their web services (see here), and hopefully get people to build cool stuff using government data (see Data.gov for more). See here for a nice list of the US government’s web services. One of these agencies was the United States Geological Survey (USGS). They opened up an app competition and [we won best overall app!
Real use cases from people using our software are awesome. They are important for many reasons: 1) They make the code more useable because we may change code to make the interace and output easier to understand; 2) They may highlight bugs in our code;
Scholarly metadata - the meta-information surrounding articles - can be super useful. Although metadata does not contain the full content of articles, it contains a lot of useful information, including title, authors, abstract, URL to the article, etc. One of the largest sources of metadata is provided via the Open Archives Initiative Protocol for Metadata Harvesting or OAI-PMH.

At rOpenSci we’re very passionate about engaging with our community and getting more people on board with open science and open data. There are many challenges to be overcome before this practice becomes mainstream. Even when researchers see the value in engaging more openly, the learning curve associated with various aspects of the workflow can seem daunting.
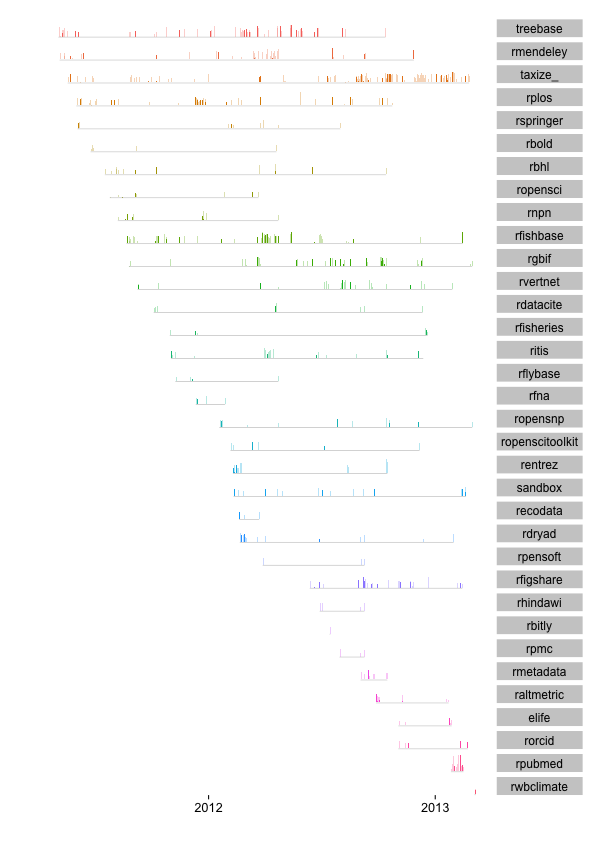
We have been writing code for R packages for a couple years, so it is time to take a look back at the data. What data you ask? The commits data from GitHub ~ data that records who did what and when. Using the Github commits API we can gather data on who commited code to a Github repository, and when they did it. Then we can visualize this hitorical record.

The following was a guest post from Ignasi Bartomeus, originally posted on his blog on 26 Nov, 2012. Check out a related blog post here. Note the functionality discussed in this post is now in our taxize package under the function gisd_isinvasive. We hacked out a quick Shiny app so you can play around with the below function in taxize on the web to get invasive status and plot it on a phylogeny. Check it out here.