
One reason I built BioNames (and the related digital archive BioStor) was to create tools to help make sense of taxonomic names.

One reason I built BioNames (and the related digital archive BioStor) was to create tools to help make sense of taxonomic names.

Bob Mesibov (who has been a guest author on this blog) recently published a paper on data quality in in ZooKeys :In this paper Bob documents some significant discrepancies between data in his Millipedes of Australia (MoA) database and the equivalent data in the Atlas of Living Australia and GBIF (disclosure, I was a reviewer of the paper, and also sit on GBIF's science committee). This paper spawned a thread on TAXACOM, and also came up
Things are finally coming together, at least enough to have a functioning demo. It looks awful, but shows the main things I want BioNames to do. One thing I'm most concerned about at this stage is the possible confusion users might experience between taxon names and concepts.
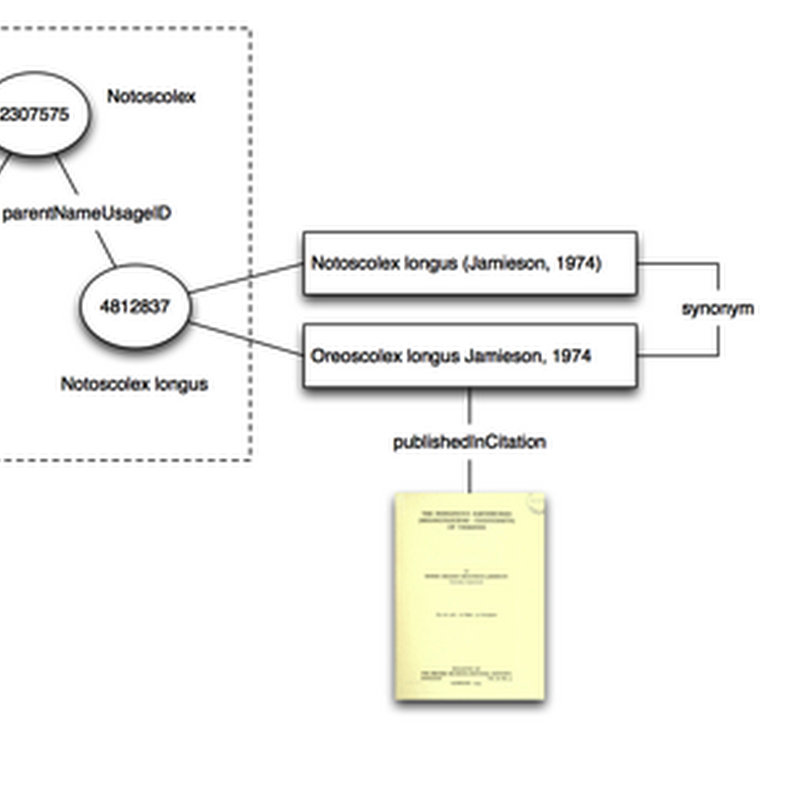
Quick notes on "taxon concepts". In order to navigate through taxon names I plan to have at least one taxonomic classification in BioNames. GBIF makes the most sense at this stage. The model I'm adopting is that the classification is a graph where nodes have the id used by the external database (in this case GBIF). Each node has one or more names attached, and where possible the names are linked to the original description.
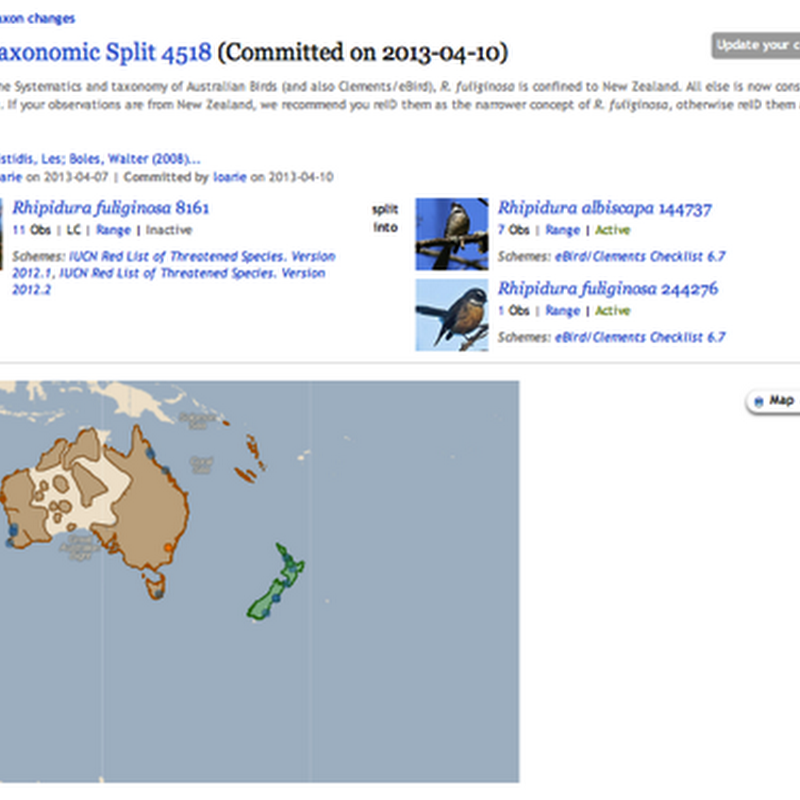
Donald Hobern drew my attention to nice the way iNaturalist displays taxonomic splits: In this example, observations identified as Rhipidura fuliginosa are being split into Rhipidura fuliginosa and Rhipidura albiscapa . This immediately reminds me of the idea which keeps circulating around, namely using version control tools to manage taxonomic classification.
On eof the things BioNames will need to do is match taxon names to classifications. For example, if I want to display a taxonomic hierarchy for the user to browse through the names, then I need a map between the taxon names that I've collected and one or more classifications. The approach I'm taking is to match strings, wherever possible using both the name and taxon authority.

Somehow I get the feeling that botanists haven't got the "open data" religion. Not only is the list of plant names list behind a really bad license, but the Global Plants Initiative (GPI) hides its type images behind a JSTOR Plant Sciences paywall.
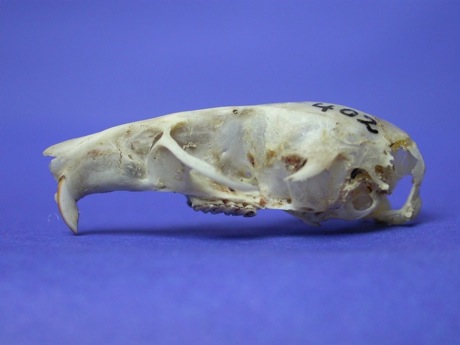
Continuing the theme of trying to map specimens cited in the literature to the equivalent GBIF records, consider the GBIF record http://data.gbif.org/occurrences/685591320, which according to GBIF is specimen "ZFMK 188762" (a [sic] holotype of Praomys hartwigi ).This is odd, because the original publication of this name (Eisentraut, M. 1968 .Beitrag zur Saugetierfauna von Kamerun.

We all have a "past" that we might not advertise widely, and my past includes flirting with panbiogeography.
If we are ever going to link biodiversity data together we need to have some way of ensuring persistent links between digital records. This isn't going to happen unless people take persistent identifiers seriously.I've been trying to link specimen codes in publications to GBIF, with some success, so imagine my horror when it started to fall apart.