
Quick notes on yet another attempt to marry the task of editing a taxonomic classification with versioning it in GitHub.The idea of dumping the whole GBIF classification into GitHub as a series of nested folders looks untenable.

Quick notes on yet another attempt to marry the task of editing a taxonomic classification with versioning it in GitHub.The idea of dumping the whole GBIF classification into GitHub as a series of nested folders looks untenable.
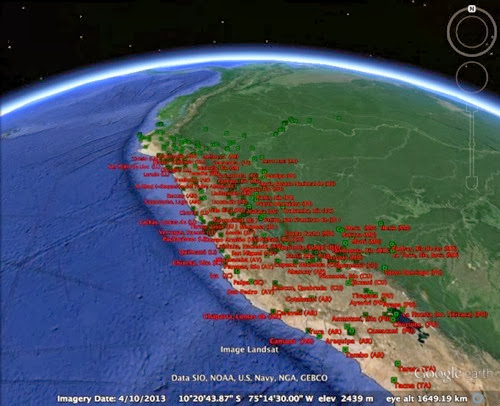
Here's another example of a Darwin Core Archive that is "broken" such that GBIF is missing some information. GBIF data set A checklist to the wasps of Peru (Hymenoptera, Aculeata) comes from Pensoft, and corresponds to the paper:As with the previous example GBIF says there are 0 georeferenced records in this dataset. This is odd, because the ZooKeys page for this article lists three supplementary files, including KML files for Google Earth.

Following on from Annotating and cleaning GBIF data: Darwin Core Archive, GitHub, ORCID, and DataCite here's a quick and dirty example of using GitHub to help clean up a Darwin Core Archive. The dataset 3i - Cicadellinae Database has 2,152 species and 4,749 taxa, but GBIF says it has no georeferenced data.

This is a quick sketch of a way to combine existing tools to help clean and annotate data in GBIF, particularly (but not exclusively) occurrence data. GitHub The data provider puts a Darwin Core Archive (expanded, not zipped) into a GitHub repository.
I've recently been appointed Chair of the Science Committee of the Global Biodiversity Information Facility (GBIF) http://www.gbif.org [1]. The committee is a small group of people with a range of backgrounds, and one of our roles is to advise GBIF on matters scientific (e.g., what kinds of data GBIF should collect?, what kinds of scientific questions should GBIF help answer?, etc.).There have been formal surveys (see the papers in the journal

Wednesday saw the launch of the Global Biodiversity Informatics Outlook (GBIO), based in large part on the Global Biodiversity Informatics Conference (GBIC). The aim is to provide a framework for biodiversity informatics and its applications in the hope that the field will unite around a shared vision of where we are and what needs to be done next:There is a web site http://www.biodiversityinformatics.org/ with more details and links to related
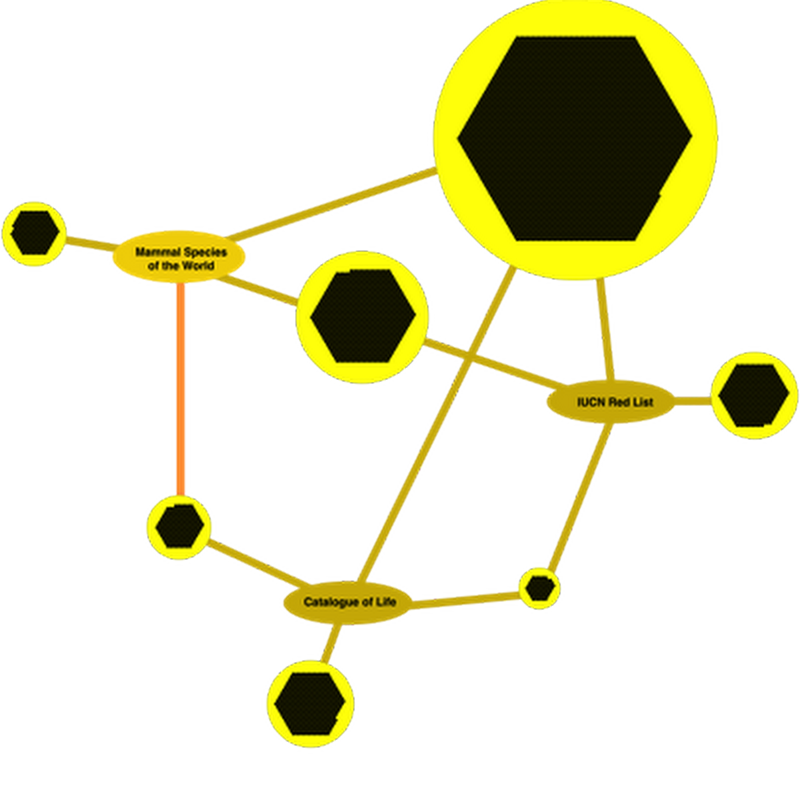
In some recent posts I've been exploring the quality of GBIF's taxonomic data. I've done some further analyses and decided to write this up in something more than a blog post. I'm writing a draft which you can see on GitHub. It tackles just one issue, namely what happens when you combine taxonomic names from multiple sources and don't know that some of those names are synonyms.

Continuing the theme of the failings of the GBIF classification I've been playing further with cluster maps to visualise the problem (see this earlier post for an introduction).Browsing through bats in GBIF I keep finding the same species appearing more than once, albeit in different genera.
GBIF is asking for views on how it should license of data in the GBIF network. The full consultation document is available from Google Drive and DropBox.
In browsing the GBIF classification in BioNames I keep coming across cases of wholesale duplication of taxa.