Things are finally coming together, at least enough to have a functioning demo. It looks awful, but shows the main things I want BioNames to do. One thing I'm most concerned about at this stage is the possible confusion users might experience between taxon names and concepts.
iPhylo

Over on Google Plus (yeah, me neither) Donat Agosti is giving me a hard time regarding the quality of some data that I am using.

This seems to be the season for big, arm-wavy documents about the future of biodiversity informatics (see A decadal view of biodiversity informatics: challenges and priorities). An equivalent document is being drafted based on the Global Biodiversity Informatics Conference (GBIC 2012) conference.

In an earlier post I discussed using Open Refine (formerly Google Refine) to clean and reconcile taxon names. I've added an additional service that can be used to reconcile author names that uses the Virtual International Authority File (VIAF) API.

BMC Ecology has published Alex Hardisty and Dave Roberts' white paper on biodiversity informatics: Here are their 12 recommendations (with some comments of my own): Open Data, should be normal practice and should embody the principles of being accessible, assessable, intelligible and usable. Seems obvious, but data providers are often reluctant to open "their" data up for reuse. Data encoding should allow analysis across multiple
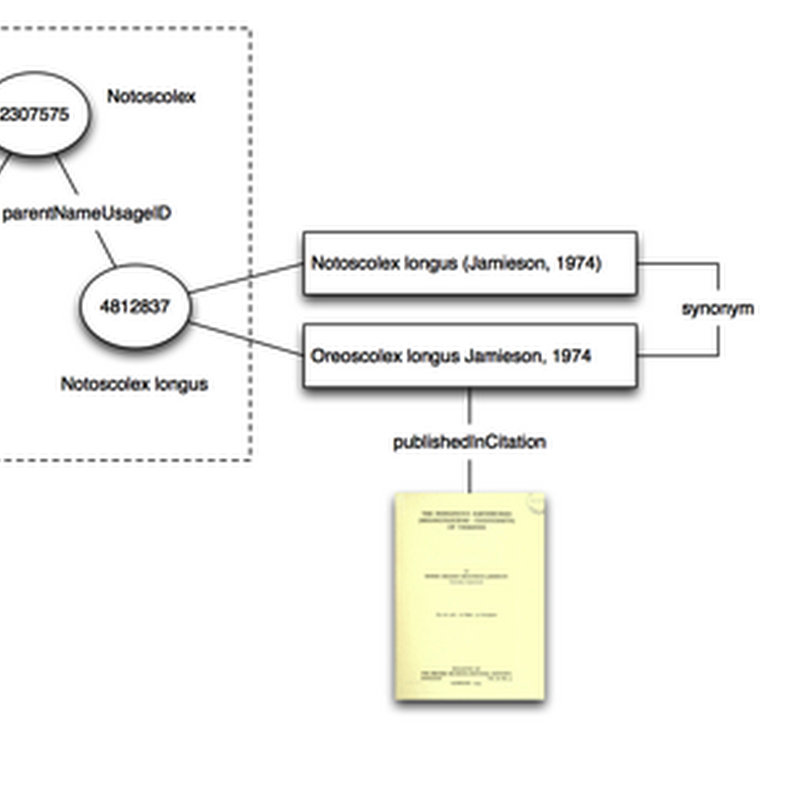
Quick notes on "taxon concepts". In order to navigate through taxon names I plan to have at least one taxonomic classification in BioNames. GBIF makes the most sense at this stage. The model I'm adopting is that the classification is a graph where nodes have the id used by the external database (in this case GBIF). Each node has one or more names attached, and where possible the names are linked to the original description.
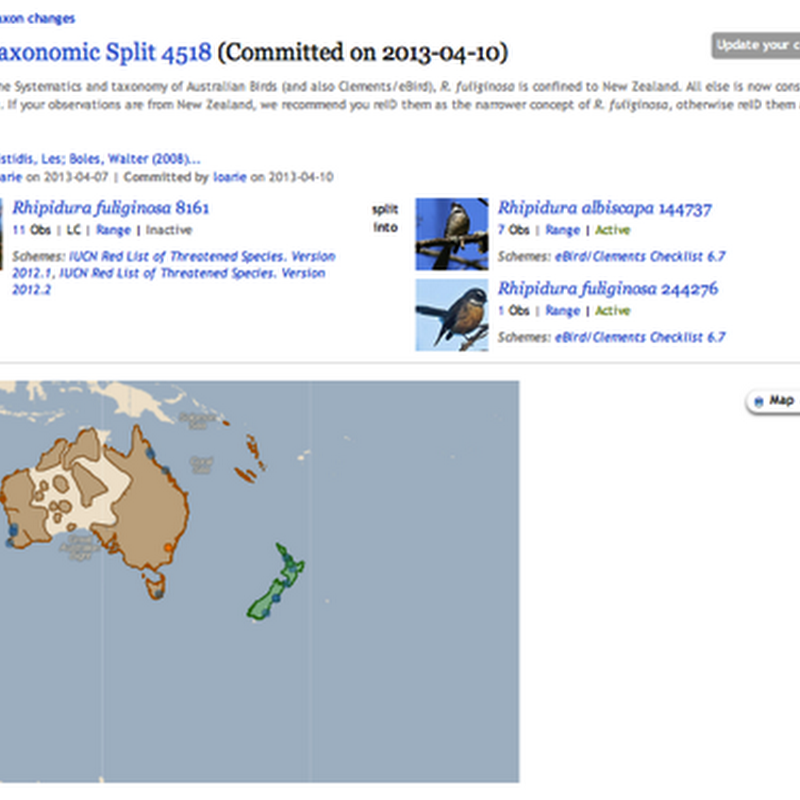
Donald Hobern drew my attention to nice the way iNaturalist displays taxonomic splits: In this example, observations identified as Rhipidura fuliginosa are being split into Rhipidura fuliginosa and Rhipidura albiscapa . This immediately reminds me of the idea which keeps circulating around, namely using version control tools to manage taxonomic classification.

Came across this paper recently:Despite QR Codes being uncool, there's something appealing about the idea of compressing a DNA barcode sequence into a small image.
I'm working on displaying OCR text from BHL using SVG, and these are just some quick notes on font size. Specifically how SVG font size corresponds to the size of letters, and how you work out what point size was used to print text on a BHL page.SVG font-size corresponds to the EM square of the font.

The new look Biodiversity Heritage Library includes articles extracted from BioStor, which is a step forwards in making the "legacy" biodiversity literature more accessible. But we still have some way to go. In particular the articles lack the obvious decoration of a modern article, the DOI.