Update: Angelique Hjarding and her co-authors have responded in a guest post on iPhylo.The quality and fitness for use of GBIF-mobilised data is a topic of interest to anyone that uses GBIF data.
iPhylo
I stumbled across this paper (found on the GBIF Public Library):The first sentence of the abstract makes the paper sound a bit of a slog to read, but actually it's a great fun, full of pithy comments on the state of digital humanities. Almost all of this is highly relevant to mobilising natural history data.

I've been involved in a few Twitter exchanges about the upcoming pro-iBiosphere meeting regarding the "Open Biodiversity Knowledge Management System (OBKMS)", which is the topic of the meeting.
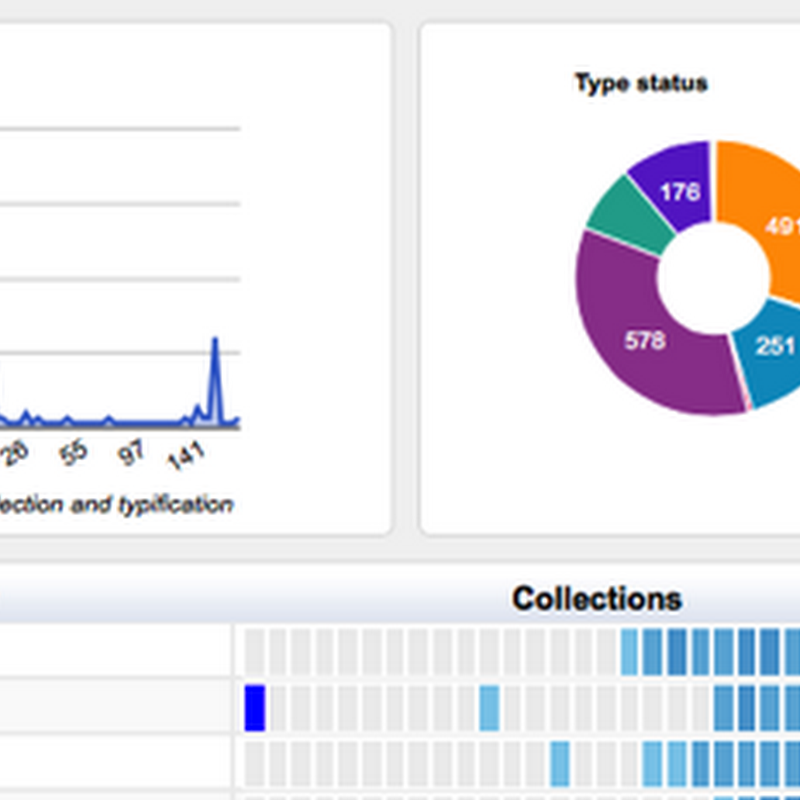
I'm adding more charts to the GBIF Chart tool, including some to explore the type status of specimens from the Solomon Islands.

Note to self on citation matching. Looking for this paper "Fishes of the Marshall and Marianas islands. Vol. I. Families from Asymmetrontidae through Siganidae" I Googled it, adding "bistro" as a search term to see if I'd already added it to BioStor.
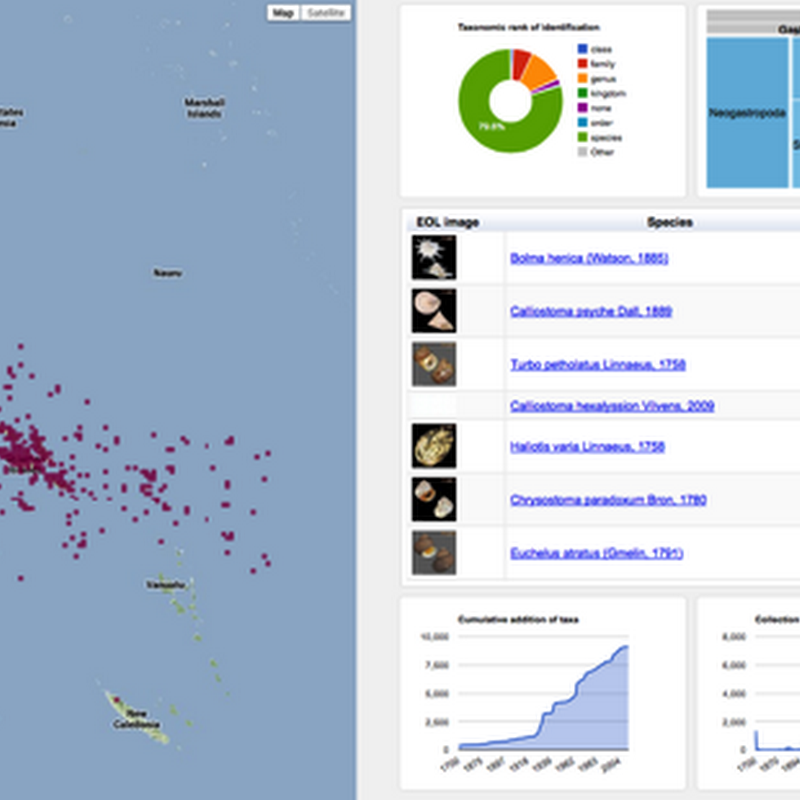
Following on from the previous post on visualising GBIF data, I've added some more interactivity.
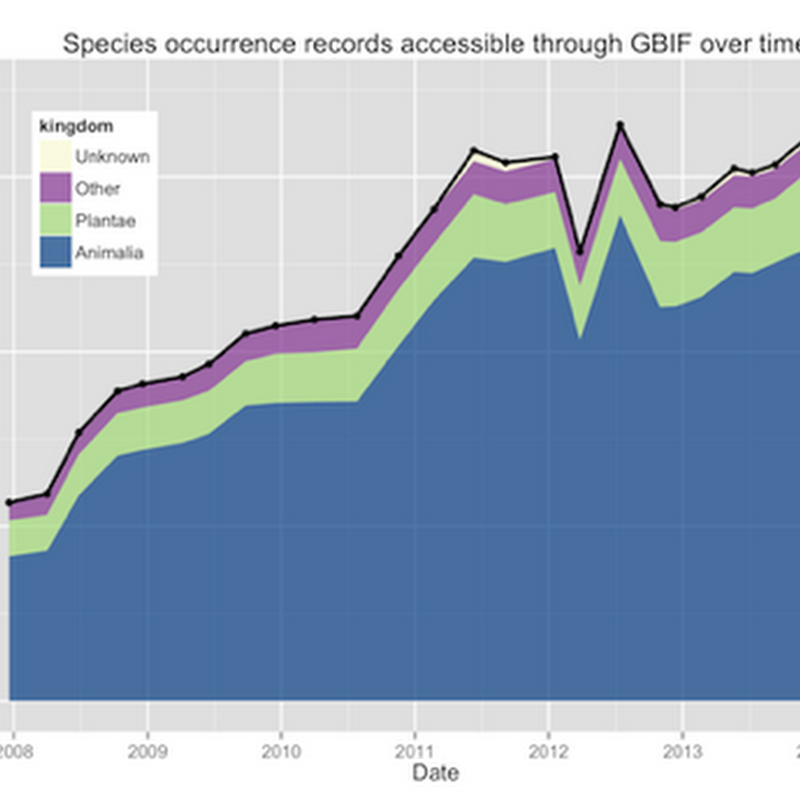
Tim Roberston and the ream at GBIF are working on some nice visualisations of GBIF data, and have made an early release available for viewing: http://analytics.gbif-uat.org.

It is almost a year to the day that I released BioNames, a database of "taxa, texts, and trees". This project was my entry in EOL's Computable Data Challenge.
I had a long Twitter conversation with Terry Catapano (@catapanoth) today, and as can happen with a distracted stream of tweets, I think we were a little at cross purposes. This blog post is an attempt to unpack the debate.
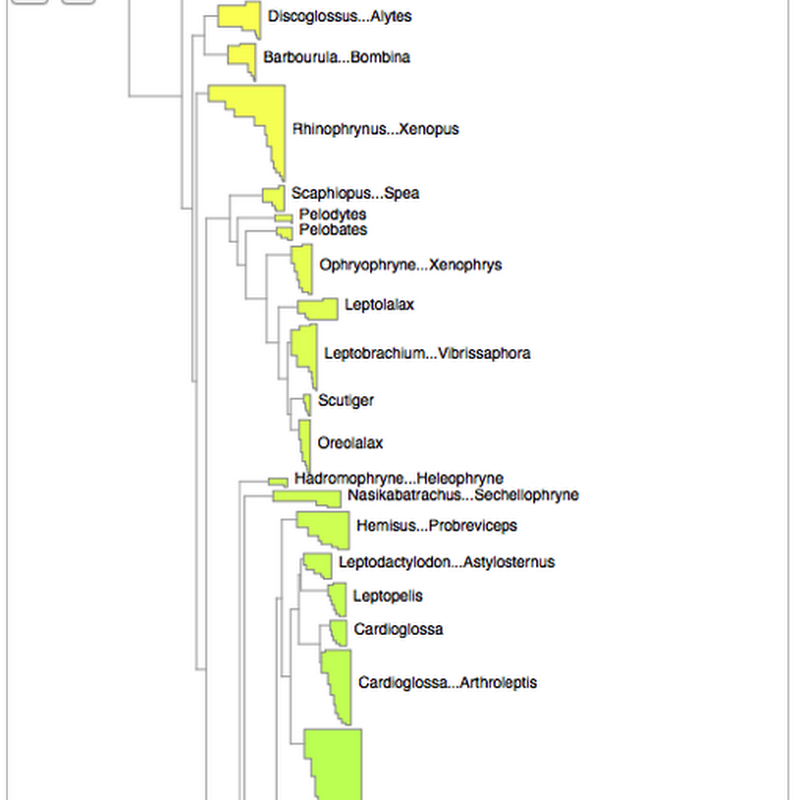
As announced on phylobabble I've started to revisit visualising large phylogenies, building on some work I did a couple of years ago (my how time flies). This time, there is actual code (see https://github.com/rdmpage/deep-tree) as well as a live demo http://iphylo.org/~rpage/deep-tree/demo/. You can see the amphibian tree below at http://iphylo.org/~rpage/deep-tree/demo/show.php?id=5369171e32b7a:You can upload or paste a tree (for now in NEXUS