
Duplicate records are the bane of any project that aggregates data from multiple sources.

Duplicate records are the bane of any project that aggregates data from multiple sources.
This message appeared on the TAXACOM mailing list:Given that most specimens lack resolvable digital identifiers (a theme I've harped on about before, most recently in the context of DNA barcoding), answering this kind of query ends up being a case of searching publications for text strings that contain the acronym of the collection.

Given various discussions about identifiers, dark taxa, and DNA barcoding that have been swirling around the last few weeks, there's one notion that is starting to bug me more and more.
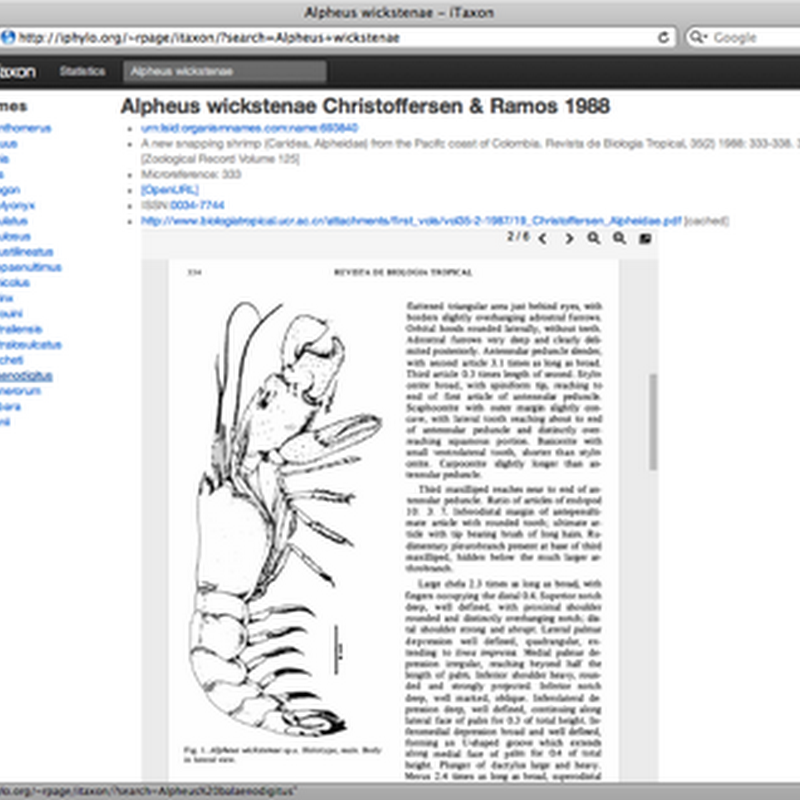
Following on from my earlier post Linking taxonomic names to literature: beyond digitised 5×3 index cards I've been slowly updating my latest toy:http://iphylo.org/~rpage/itaxonThis site displays a database mapping over 200,000 animal names to the primary literature, using a mix of identifiers (DOIs, Handles, PubMed, URLs) as well as links to freely available PDFs where they are available.

Geoffery Bilder's comments about the unsuitability of URLs as long term identifiers (as opposed, say, to DOIs) came to mind when I discovered that the domain phthiraptera.org is up for sale: This domain used to be home to a wealth of resources on lice (order Phthiraptera). I discovered that ownership of the domain had expired when a bunch of links to PDFs returned by an iSpecies search for Collodennyus all bounced to the holding page

I've been playing recently with the Biodiversity Heritage Library (BHL), and am starting to get a sense for the complexities (and limitations) of the metadata BHL stores about publications.

Continuing with RSS feeds, I've now added wrappers around IPNI that will return for each plant family a list of names added to the IPNI database in the last 30 days. You can see the list at here.One thing which is a constant source of frustration for me is the disconnect between nomenclators (lists of published names for species) and scientific publishing.

Duncan Hull alerted me to his paper "Defrosting the Digital Library: Bibliographic Tools for the Next Generation Web" ( PloS Computational Biology , doi:10.1371/journal.pcbi.1000204). Here's the abstract:It's an interesting read, and it also cites my bioGUID project.[Image from dave 7]

I've been using ISSN's (International Standard Serial Number) to uniquely identify journals, both to generate article identifiers, and as a parameter to send to CrossRef's OpenURL resolver. Recently I've come across journals that change their ISSN, which has fairly catastrophic effects on my lookup tools.

As much as I like the idea of a globally unique, resolvable identifier, my recent experience with JSTOR is making me wonder.JSTOR has three identifiers for articles it archives, DOIs, SICIs, and stable URLs (the later being introduced with the new platform released April 4, 2008). Previously JSTOR would publish DOIs for many of its articles.