
There is a great post by Jeni Tennison on the Open Data Institute blog entitled Five Stages of Data Grief.

There is a great post by Jeni Tennison on the Open Data Institute blog entitled Five Stages of Data Grief.

I gave a remote presentation at a proiBioSphere workshop this morning. The slides are below (to try and make it a bit more engaging than a desk of Powerpoints I played around with Prezi).There is a version on Vimeo that has audio as well.I sketched out the biodiversity "knowledge graph", then talked about how mark-up relates to this, finishing with a few questions.

Scott Federhen told me about a nice new feature in GenBank that he's described in a piece for NCBI News. The NCBI taxonomy database now shows a its of type material (where known), and the GenBank sequence database "knows: about types. Here's the summary:You can query for sequences from type using the query "sequence from type"[filter]. This could lead to some nice automated tools.

VertNet has announced that they have implemented issue tracking using GitHub. This is a really interesting development, as figuring out how to capture and make use of annotations in biodiversity databases is a problem that's attracting a lot of attention.
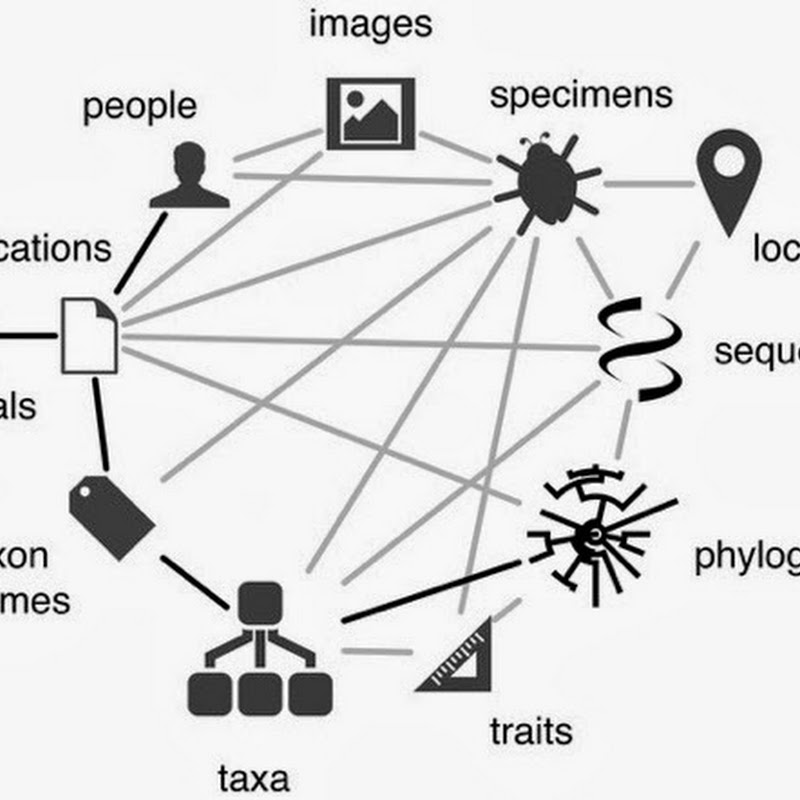
More for my own benefit than anything else I've decided to list some of the things I plan to work on this year. If nothing else, it may make sobering reading this time next year. A knowledge graph for biodiversity Google's introduction of the "knowledge graph" gives us a happy phrase to use when talking about linking stuff together. It doesn't come with all the baggage of the "semantic web", or the ambiguity of "knowledge base".
Given that it's the start of a new year, and I have a short window before teaching kicks off in earnest (and I have to revise my phyloinformatics course) I'm playing with a few GBIF-related ideas. One topic which comes up a lot is annotating and correcting errors. There has been some work in this area [1][2] bit it strikes me as somewhat complicated.

The following is a guest blog post by David Schindel and colleagues and is a response to the paper by Antonio Marques et al. in Science doi:10.1126/science.341.6152.1341-a.Marques, Maronna and Collins (1) rightly call on the biodiversity research community to include latitude/longitude data in database and published records of natural history specimens.

A while ago I posted BHL to PDF workflow which was a sketch of a work flow to generate clean, searchable PDFs from Biodiversity Heritage Library (BHL) content: I've made some progress on putting this together, as well as expanded the goal somewhat. In fact, there are several goals:BioStor articles need to be archived somewhere. At the moment they live on my server, and metadata is also served by BHL (as the "parts" you see in a scanned volume).

Quick notes on yet another attempt to marry the task of editing a taxonomic classification with versioning it in GitHub.The idea of dumping the whole GBIF classification into GitHub as a series of nested folders looks untenable.

There is a fairly scathing editorial in Nature [The new zoo. (2013). Nature, 503(7476), 311–312. doi:10.1038/503311b ] that reacts to a recent paper by Dubois et al.:To quote the editorial:Ouch! But Dubois et al.'s paper pretty much deserves this reaction - it's a reactionary rant that is breathtaking in it's lack of perspective.