
A decade ago (OMG, that can't be right, an actual decade ago) I created "iSpecies", a simple little tool to mashup a variety of data from GBIF, NCBI, Yahoo, Wikipedia, and Google Scholar to create a search engine for species.

A decade ago (OMG, that can't be right, an actual decade ago) I created "iSpecies", a simple little tool to mashup a variety of data from GBIF, NCBI, Yahoo, Wikipedia, and Google Scholar to create a search engine for species.
My paper "Surfacing the deep data of taxonomy" (based on a presentation I gave in 2011) has appeared in print as part to a special issue of Zookeys : The manuscript was written shortly after the talk, but as is the nature of edited volumes it's taken a while to appear. My tweet about the paper sparked some interesting comments from David Shorthouse.

OK, so the title is pure click bait, but here's the thing. It seems to me that the Semantic Web as classically conceived (RDF/XML, SPARQL, triple stores) has had relatively little impact outside academia, whereas other technologies such as JSON, NoSQL (e.g., MongoDB, CouchDB) and graph databases (e.g., Neo4J) have got a lot of developer mindshare. In biodiversity informatics the Semantic Web has been a round for a while.

It's a nice feeling when work that one did ages ago seems relevant again. Markus Döring has been working on a new backbone classification of all the species which occur in taxonomic checklists harvested by GBIF. After building a new classification the obvious question arises "how does this compare to the previous GBIF classification?" A simple question, answering it however is a little tricky.
For those of you who, like me, weren't at the "Frontiers Of Biodiversity Informatics and Modelling Species Distributions" held at the AMNH in New York, here are the videos of the talks and panel discussion, which the organisers have kindly put up on Vimeo with the following description:

This guest post by Tony Rees explores some of the themes from his recent talk 10 years of Global Biodiversity Databases: Are We There Yet?. 10 years of global biodiversity databases: are we there yet?
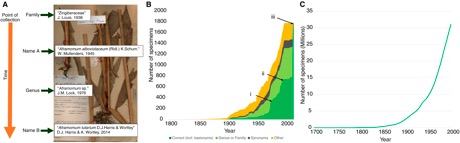
Zoë A. Goodwin (@Drypetes) and collegagues have published a paper with a title guaranteed to get noticed: Their paper argues that "more than half of all tropical plant collections may be wrongly named." This is clearly a worrying conclusion with major implications for aggregators such as GBIF that get the bulk of their data (excluding birds) from museums and herbaria.
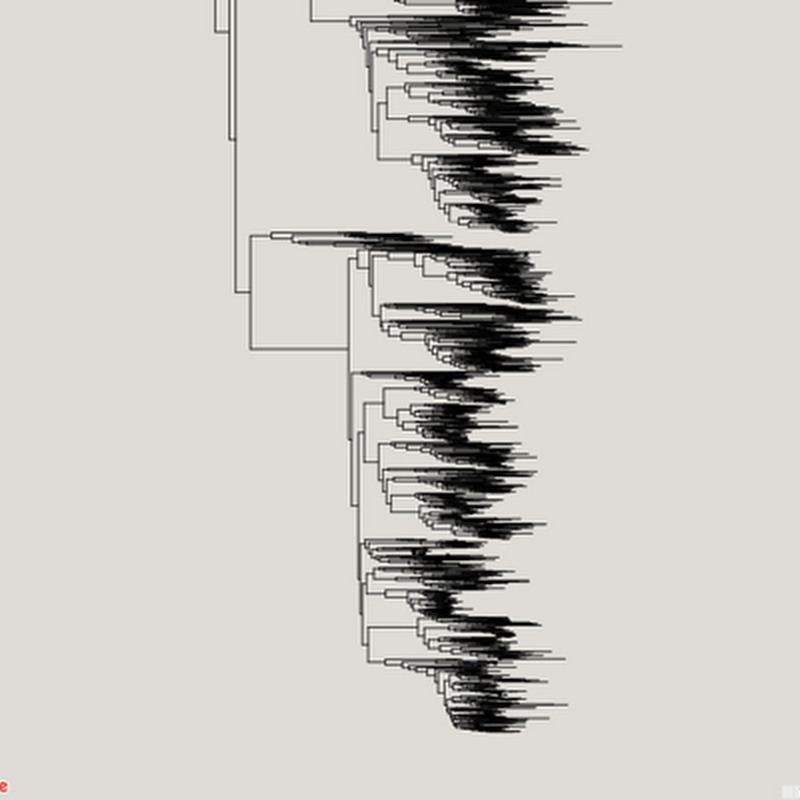
Inspired in part by the release of the draft tree of life (doi:10.1073/pnas.1423041112 by the Open Tree of Life, I've been revisiting (yet again) ways to visualise very big phylogenies (see Very large phylogeny viewer for my last attempt). My latest experiment uses Google Maps to render a large tree. Google Maps uses "tiles" to create a zoomable interface, so we need to create tiles for different zoom levels for the phylogeny.

Currently in classes where I teach the basics of tree building, we still fire up ancient iMacs, load up MacClade, and let the students have a play. Typically we give them the same data set and have a class competition to see which group can get the shortest tree by manually rearranging the branches. It’s fun, but the computers are old, and what’s nostalgic for me seems alien to the iPhone generation.

On Friday I discovered that BHL has started issuing CrossRef DOIs for articles, starting with the journal Revue Suisse de Zoologie . The metadata for these articles comes from BioStor. After a WTF and WWIC moment, I tweeted about this, and something of a Twitter storm (and email storm) ensued: To be clear, I'm very happy that BHL is finally assigning article-level DOIs, and that it is doing this via CrossRef.